Equivariant Geometric Graph Neural Networks
Equivariant Geometric Graph Neural Networks
Message Passing
Most graph neural networks are formulated under the message passing framework, where node representations are updated by exchanging information with neighboring nodes. At each layer, information is propagated along graph edges through a sequence of message construction, aggregation, and update steps.
For a given edge connecting two nodes, it is convenient to define a direction for information flow. Although molecular graphs are typically undirected, message passing is implemented as two directed operations per edge to simplify notation and computation. In this formulation, one node generates a message based on its current state, which is then sent to its neighbor and aggregated with other incoming messages.
We can consider node j as the message sender, while node i is the message receiver.
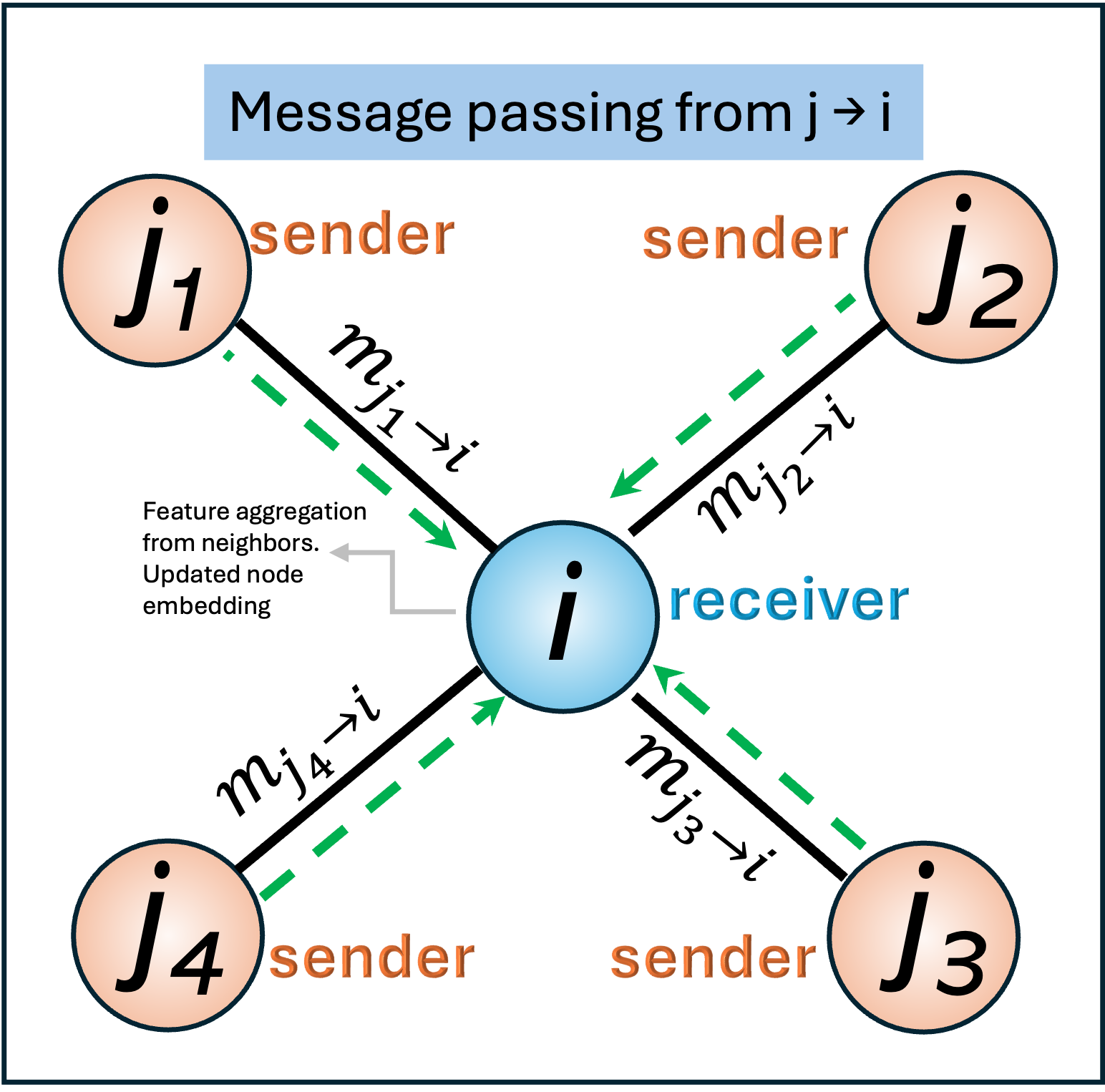
This sender–receiver convention allows GNNs to model asymmetric interactions, incorporate edge features, and naturally extend to directed graphs and attention-based mechanisms. In physical systems, this perspective aligns with the idea that each atom contributes an interaction-dependent influence to its neighbors, which are then combined to determine the local environment of the receiving atom.
Equivariance vs Invariance
Invariance: A function is invariant if its output does not change when the input is transformed.
“Move or rotate the molecule/protein — the prediction stays the same.”
Example:
- Molecular energy
- Binding affinity
- Toxicity score
Equivariance: A function is equivariant if its output transforms in the same way as the input.
“Move or rotate the molecule/protein — the output moves or rotates in exactly the same way. This is associated with the group transformation (group theory).”
Example:
- Atomic forces
- Velocity vectors
- Dipole moments
In molecular and protein systems, physical laws are independent of the choice of coordinate system. If a molecule is translated or rotated in space, its energy, binding affinity, and chemical identity do not change. Forces and velocities, however, must rotate consistently with the molecule.
Equivariant geometric GNNs encode this prior directly into the model architecture, ensuring that predictions transform correctly under Euclidean symmetries.
Group Theory
Euclidean Groups
- E(n): translations, rotations, and reflections in \(\mathbb{R}^n\)
- SE(3): translations + rotations (no reflections)
A function \(f\) is equivariant to group \(G\) if: \[ f(g \cdot x) = g \cdot f(x), \quad \forall g \in G \]
| Quantity | Transformation under rotation |
|---|---|
| Atomic coordinates | Rotate |
| Interatomic distances | Invariant |
| Energy | Invariant |
| Forces | Rotate |
| Dipole moments | Rotate |
Traditional GNNs often enforce invariance only. Equivariant GNNs preserve the full transformation structure.
Design Principles
Equivariant models rely on three key ideas:
- Relative geometry only
- Use \(x_i - x_j\), not absolute positions
- Scalar nonlinearities
- Apply nonlinear functions only to invariant quantities
- Structured feature types
- Distinguish scalars, vectors, and higher-order tensors
These constraints guarantee equivariance by construction.
EGNN
Philosophy
E(n) Equivariant Graph Neural Networks (EGNN) aims to provide the simplest possible equivariant GNN:
- No spherical harmonics
- No explicit group representations
- Minimal computational overhead
Mathematical Structure
Each node has:
- Features \(h_i \in \mathbb{R}^d\)
- Coordinates \(x_i \in \mathbb{R}^n\)
Messages depend only on invariant quantities: \[ m_{ij} = \phi_e(h_i^{l}, h_j^{l}, \|x_i^{l} - x_j^{l}\|^2, a_{ij}) \]
- \(h_i^{l}, h_j^{l}\): The attributes of atoms/residues \(i\), \(j\), respectively;
- \(\\|x_i^{l} - x_j^{l}\\|^2\): The relative squared distance between two coordinates/atoms/residues;
- \(a_{ij}\): The edge/bond/interaction attributes.
Coordinates are updated using scaled relative displacements: \[ x_i^{l+1} = x_i^{l} + C\sum_j (x_i^{l} - x_j^{l}) \, \phi_x(m_{ij}) \]
- \(x_i^{l+1}\): Coordinates of residue/atom \(i\) at layer \(l+1\)
- \(x_i^{l}\): Coordinates of residue/atom \(i\) at (previous) layer \(l\)
- \(x_i^{l}\) \(x_j^{l}\): Coordinates of residue/atom \(i\) and \(j\) at layer \(l\)
Nodes are updated based on the aggregation of messages \[ m_i = \sum_j (m_{ij}) \]
\[ h_i^{l+1} = \phi_h(h_i^{l}, m_i) \]
Why This Is Equivariant
- Relative displacement \(x_i - x_j\) rotates under \(R\)
- Scalar \(\phi_x(m_{ij})\) does not
- Product rotates correctly
- Summation preserves equivariance
Strengths
- Extremely simple
- Fast and stable
- Easy to integrate into existing GNN pipelines
Weaknesses
- Limited expressivity for directional or angular interactions
- Cannot represent higher-order tensor features
Typical Use Cases
- Molecular dynamics
- Protein structure refinement
- 3D point cloud regression
- Fast physics surrogates
SE(3)-Transformer
Philosophy
SE(3)-Transformers pursue maximum expressivity, explicitly modeling the representation theory of the rotation group.
They treat node features as irreducible representations (irreps):
- Scalars (\(l=0\))
- Vectors (\(l=1\))
- Higher-order tensors (\(l \ge 2\))
Equivariant Attention
Messages are constructed using:
- Attention weights (scalars)
- Radial functions of distance
- Spherical harmonics \(Y_{l,m}(\hat{r}_{ij})\)
Conceptually: \[ m_{ij}^{(l)} = \sum_{l’} \alpha_{ij} \, R_{l,l’}(r_{ij}) \, Y_{l,m}(\hat{r}_{ij}) \, h_j^{(l’)} \]
Why This Is Powerful
- Encodes angles and directions explicitly
- Preserves equivariance at every layer
- Can model anisotropic interactions
Strengths
- Very high expressivity
- Strong performance on structure-sensitive tasks
- Suitable for protein and materials modeling
Weaknesses
- Complex implementation
- High memory and compute cost
- Requires specialized libraries (e3nn)
Typical Use Cases
- Protein structure prediction
- Binding site modeling
- High-accuracy force fields
EGNN vs SE(3)-Transformer
| Aspect | EGNN | SE(3)-Transformer |
|---|---|---|
| Equivariance | E(n) | SE(3) |
| Feature types | Scalars only | Scalars + vectors + tensors |
| Angular info | Implicit (limited) | Explicit (via spherical harmonics) |
| Complexity | Low | High |
| Speed | Fast | Slow |
| Expressivity | Moderate | Very high |
| Implementation | Simple PyTorch | Requires equivariant ops |
Relation to Physics-Based Modeling
Equivariant GNNs bridge ML and physics by:
- Respecting symmetry laws
- Reducing sample complexity
- Improving out-of-distribution generalization
They are often used as:
- Surrogate force fields
- Differentiable simulators
- Geometry-aware generative models
In practice, they complement rather than replace molecular dynamics and quantum chemistry.
Practical Guidelines for Drug Discovery
- No 3D structure? → topology-based GNN
- 3D structure, invariant target? → SchNet
- Need forces or coordinate updates? → EGNN
- Need directional accuracy? → SE(3)-Transformer
A common workflow:
Equivariant GNN → candidate generation → physics-based refinement (MD / docking)
Key References
- Satorras et al., E(n)-Equivariant Graph Neural Networks, ICML 2021
- Fuchs et al., SE(3)-Transformers, NeurIPS 2020
- Batzner et al., E(3)-Equivariant Graph Neural Networks, NeurIPS 2022
- Thomas et al., Tensor Field Networks, NeurIPS 2018
Enjoy Reading This Article?
Here are some more articles you might like to read next: